Hyperspectral Dark-Field Microscopy Enables Rapid and Accurate Identification of Cancerous Tissues
May 15, 2024
Source: drugdu
 658
658
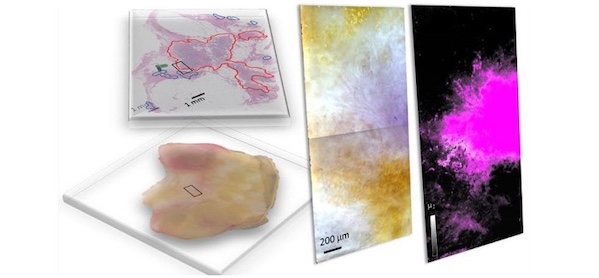
Breast cancer remains a major cause of cancer-related mortality among women. Breast-conserving surgery (BCS), also known as lumpectomy, is the removal of the cancerous lump and a small margin of surrounding tissue. This procedure is typically advised for women with early-stage breast cancer or small tumors, as it conserves more of the breast tissue compared to a mastectomy. After undergoing BCS, it is critical to verify that all cancerous cells have been removed to decide if additional surgery is necessary. This verification involves a tumor margin assessment, which examines the edges of the excised tissue (tumor margins) to check for residual cancer cells. Conventionally, this assessment entails staining the tissue samples with dyes and inspecting them under a microscope to differentiate between healthy and cancer cells. However, new optical imaging techniques have emerged as quicker alternatives for conducting these assessments.
A group of researchers from the United States, including members from the National Institute of Standards and Technology (NIST, Gaithersburg, MD, USA), has introduced hyperspectral dark-field microscopy (HSDFM) as an effective technique to swiftly and accurately distinguish between cancerous and healthy cells and identify various tumor subtypes in breast tissues post-lumpectomy. In HSDFM, tissue samples are exposed to multiple wavelengths of light, and the varying intensity of light scattered by cellular and molecular components is analyzed to create distinctive spectral signatures for each type of tissue. This technique generates two-dimensional images where each pixel holds spectral data across multiple wavelengths, enabling precise identification of tissue composition. This approach specifically tackles the limitations commonly faced in hyperspectral tumor margin imaging techniques, which typically depend on reflectance to collect spectral information from tissue samples.
Reflectance-based imaging techniques often struggle with issues like the uneven absorption of light by biological substances, such as oxyhemoglobin in blood, which can lead to inconsistent spectral signatures from different samples. In their study, the researchers examined HSDFM images of breast lumpectomy specimens from several patients. They employed two machine learning strategies to categorize the pixels by tissue type: a supervised method and an unsupervised method. The supervised method utilized was spectral angle mapping, which involves comparing the spectral signature of each pixel against known spectral signatures of different tumor subtypes and tissue types (like fat, connective tissue, and blood) previously identified via histopathological analysis.
For the unsupervised method, they applied the K-means clustering algorithm, which sorts pixels into clusters based on similarity in their spectral signatures, thereby aiding in the identification of tumor regions without needing prior spectral data or specific tissue type knowledge. The spectral signatures derived from both the supervised and unsupervised methods were similar and effectively pinpointed areas containing invasive ductal carcinoma—the most prevalent form of breast cancer, accounting for 75% of all cases—as well as invasive mucinous carcinoma, a less common type where cancer cells grow in mucus. The results indicate that the unsupervised approach is validated by the supervised method, suggesting that HSDFM imaging data could be instrumental in developing unsupervised algorithms for the quick and accurate detection of cancerous tissues, which is expected to improve post-surgical monitoring and treatment planning in BCS, enabling more timely interventions.
Source:
https://www.labmedica.com/pathology/articles/294801131/hyperspectral-dark-field-microscopy-enables-rapid-and-accurate-identification-of-cancerous-tissues.html
Read more on
- Cinda leads the world, and Takeda’s 1.2 billion gamble is reaping the rewards. June 5, 2026
- The National Medical Products Administration (NMPA) conditionally approved the marketing of Pometamethasone Alpha for injection. June 5, 2026
- The generic version of this drug, developed by 35 pharmaceutical companies, has had its market launch delayed by 5 years. June 5, 2026
- Pharmaceutical giant lays off more employees June 5, 2026
- Imported “new kidney disease drug” approved June 5, 2026
your submission has already been received.
OK
Subscribe
Please enter a valid Email address!
Submit
The most relevant industry news & insight will be sent to you every two weeks.



